what are the types of Cancer gene?
Cancer gene
Cell division is accomplished by a series of tightly controlled processes. The process depends on the normal transcription of some genes ( Transcription ) and translation ( – Search.com ). If these processes are abnormal, it can lead to uncontrolled cell growth. It is now believed that there are approximately 30,000 genes in the human genome, and a small number of genes appear to play an important role in the prevention, occurrence and progression of tumors. In many different types of malignancies, these genes show dysfunction or loss of function.
The identified genes can be classified into the following two categories according to their normal functions in cells:
- The first type of genes are their protein products that stimulate or enhance the ability of cells to divide and survive. This class of genes also includes genes that inhibit cell death and play a role in tumor growth.
- The second type of gene is that their protein products can directly or indirectly prevent cell division or cause cell death.
The normal “version” of the first class of genes is called proto-oncogenes. However, “mutant” versions or “damaged” versions of such genes become oncogenes.
The second type of gene can be called tumor suppressors.
Please note whether the nouns related to “gene” in the text are “italic”. By convention, if the gene noun is “italic”, it means “gene” itself; if it is not “italic”, it means “protein produced by the gene”. For example, p53means “gene”; and p53 means “protein produced by this gene”.
The topics on this page are:
- cancer Gen
- HER-2 / Neu
- HER-2 / neu and cancer treatment
- RAS
- MYC
- SRC
- Telomerase
- BCL-2
- EGFR
- Oncogene form
- Summary of this section: Oncogene
- Understanding the process: oncogenes
- Tumor suppressor
- Introduction of p53
- P53 function
- Retinoblastoma gene ( Rb )
- Normal function of the Rb gene
- Introduction to APC
- APC function
- Introduction to BRCA
- BRCA features
- Table of tumor suppressor genes
- Summary of this section: Tumor suppressor gene
- MicroRNA
- MicroRNAs and cancer
Oncogenes
The role of tumor suppressor genes and oncogenes in the human body can be compared with cars. The proto-oncogene is like the car throttle in the animation below, which controls the movement of the car. When the car is in a normal state, the car can only be activated when the throttle is depressed. In normal cells, both intracellular and extracellular signals control the activity of oncogenes. In the animation below, these signals are represented by the growth factor of the “X” type and the foot of the throttle.
Defective oncogenes are like the fact that the throttle is always “on”. No signal is needed to activate these genes. The car will drive forward whether it is stepping on or not.
Apply this metaphor to cells: even if there is no signal to indicate cell division, the cells will continue to divide. There are two sets of each of our genes. For oncogenes, as long as one of the genes is defective, it will lead to the continuous division of cells.
A large number of genes have been identified as proto-oncogenes, many of which are responsible for providing positive signals leading to cell division. Some proto-oncogenes act to regulate cell death. As described in this section, oncogenes (that is, defective versions of these genes) can cause unregulated division of cells. This growth can occur even in the absence of normal growth-promoting signals, such as growth factors. A major feature of oncogene activity is that a defective version can lead to uncontrolled cell growth. This is completely different from the tumor suppressor gene, which must require both sets of genes to be defective to cause abnormal cell division.
To date, those identified proto-oncogenes have many different functions in cells. Despite their differences in normal function, mutant forms of these genes (carcinogenic versions) lead to unregulated cell division. Mutant proteins usually retain some of their original functions, but are no longer sensitive to the management signals that respond to them in their normal form. In the following, we have selected some oncogenes associated with many cancers and described them in detail.
HER -2/ Neu
HER-2 /neu (ie erb B-2) is a gene encoding human epidermal growth factor receptor type 2. There is a moderate amount of this receptor on some normal cells. As the name implies, this receptor is involved in the response of cells to growth factors. As shown in the figure below, this gene binds to growth factors under appropriate conditions to stimulate cell division.
In human breast cancer cases, up to 30% of cases have amplification of the HER-2/ neu gene. An increase in the number of copies of the HER-2/ neu gene results in increased expression of HER-2 protein on the cell surface, resulting in an accelerated rate of cell proliferation (as shown in the following figure).Itis currently believed that gene amplification affects the ability of tumor growth and spread, and also affects tumor response to treatment. Overexpression of this gene can make the tumor more aggressive, but it can also make the tumor more sensitive to some chemotherapy drugs. More information on gene amplification .
HER-2/neu and cancer treatment
The effect of overexpression of the HER-2/neu gene on the effectiveness of chemotherapy drugs is not known. Several studies have been conducted to determine the effect of HER2 protein on the effectiveness of chemotherapy drugs. A recent study was conducted in 140 patients with primary breast tumors using different concentrations of two chemotherapeutic drugs. The results showed that cells with stronger HER-2/neu expression were more susceptible to chemotherapeutic drugs than those with weaker expression, and the degree of tumor growth was more inhibited. HER-2/neu amplification appears to give tumor cells sensitivity to chemotherapeutic drugs rather than making them resistant. Chemotherapy drugs attack cells that are replicating genes, and HER-2/neu amplification leads to faster replication. Therefore, it can be concluded that due to the accelerated rate of cell division, tumor cells overexpressing HER2 will be killed more effectively. However, the true role of HER-2/neu amplification is not fully understood due to some contradictory results (ie, insensitivity to chemotherapeutic drugs) .
Other studies have shown that HER-2/neu overexpression is associated with estrogen receptor negative, low tumor cell differentiation, and decreased patient survival. Obviously, this proto-oncogene plays an important role in the occurrence of several malignant tumors, but the research in this area is far from over.
Antibody treatment with HER2
Cancer treatment has been directed against cancer cells that overexpress HER2 proteins. Genentech ‘s Herceptin® is a humanized monoclonal antibody that binds to the HER2 protein and blocks its activity, thereby preventing excessive cell proliferation. The animation below shows this process. Recently, Herceptin® has been combined with chemotherapy to treat malignant tumors with HER-2/neu gene amplification. More information about antibody therapy for cancer.
RAS
The products of the RAS gene are involved in the regulation of the kinase signaling pathway and in the regulation of gene transcription, thereby regulating cell growth and differentiation. In order to “open” this pathway, the ras protein must bind to intracellular guanosine triphosphate (GTP) molecules. In order to “close” this pathway, the RAS protein must cleave GTP molecules. Alterations in the RAS gene can alter the RAS protein, leaving the RAS protein no longer capable of cleavage of GTP molecules and release of GTP molecules. Such a change causes the signal path to be always “on”. This “on” signal causes cell growth and proliferation. Thus overexpression and expansion of ras leads to sustained cell proliferation, which is a critical step in tumorigenesis. Cell division is regulated by the balance of “positive” and “negative” signals. When ras transcription increases, too much of the gene protein accumulates in the cell; at this time, the “positive” signal of cell division is stronger than the “negative” signal.
Point mutations in the RAS gene typically result in the conversion of RAS from a proto-oncogene to an oncogene. The effects of such functional changes on cells are multifaceted because RAS is involved in many signaling pathways that control cell division and death. The anti-tumor drugs currently being developed are aimed at such RAS-dependent pathways. However, many studies have to be carried out to apply these drugs to the clinic.
The mutated RAS gene has been identified in malignancies of the following organs: pancreas (90%), colon (50%), lung (30%), thyroid (50%), bladder (6%), ovary (15%) , breast, skin, liver, kidney, certain types of leukemia. In the future, it is possible to use ras to identify certain malignancies. Pancreatic tumors have been difficult to diagnose, but identification of RAS gene mutations in the DNA of pancreatic cells excreted in the feces can help clinicians identify pancreatitis and pancreatic cancer.
MYC
The MYC protein is a transcription factor that controls the expression of several genes. Mutations in the MYC gene are found in many different malignancies , such as Burkitt’s lymphoma, B-cell leukemia, and lung cancer. The MYC oncogene family can be activated by gene rearrangement and gene amplification . Gene rearrangement involves the cleavage and rearrangement of chromosomes. This process can affect a large amount of DNA, affecting multiple genes. The movement of a gene or group of genes can occur within a chromosome or between different chromosomes. Such movements often result in changes in the genetic table and changes in cellular function.
A chromosomal translocation is a gene rearrangement, and a translocation between chromosomes 8 and 14 often results in overexpression of the MYC gene, ultimately leading to B cell lymphoma. The animation below shows the occurrence of translocation between two different chromosomes.
The amount of MYC protein in the cell is important because the balance of MYC protein activity depends on the other protein to compete with the MYC protein. Therefore, an increase in the content of either of these two proteins will disrupt this balance and thereby affect cell division.
In the video above, Dr. Gerard Evan discusses the role of MYC protein in cancer. Watch the full interview with Dr. Avon. ( Watch the full interview with Dr. Evan. )
MYC increased activity sometimes leads to programmed cell death, but this protective mechanism appears to be due to another oncogene bcl-2 exist and destruction, bcl-2 prevents myc induced apoptosis ( apoptosis ).
SRC
SRC is the first oncogene discovered . SRC was identified as a transformation reagent (carcinogenic reagent) for Rous sarcoma virus (RSV). This virus can infect chickens and other animals. RSV is a retrovirus (retrovirus ). It infects cells and “inserts” its genes into the DNA of this cell, which quickly leads to cancer. Therefore, this virus is called an acute transformed virus. When a chicken is infected, a large tumor often appears within two weeks. The researchers found that proteins from a gene in RSV can cause cells to grow in an abnormal way. Corresponding proto-oncogenes have also been found in the human genome. This human gene, if activated as an oncogene, causes cancer in a similar way.
The SRC protein is a tyrosine kinase. A kinase is an enzyme that transfers a phosphate group to a target molecule. This process is very important because the removal or addition of a phosphate group can alter the biomolecules, thereby regulating cell activity. The addition or removal of a phosphate group acts like a switch to regulate the activity of the target molecule. The src protein can change several target molecules, thereby transmitting signals to the nucleus, which contribute to the regulation of cellular activities.
At least nine SRC genes have been discovered . Since the mRNA produced by these genes has undergone different processing, at least 14 different proteins have been produced . Low levels of C-SRC are found in most cells . However, in certain types of malignancies, C-SRC is overexpressed, such as human neuroblastoma, small cell lung cancer, colon cancer, breast cancer, and rhabdomyosarcoma.
Telomerase
Due to the nature of the DNA replication process, the ends of the chromosome (called “telomeres”) become shorter during each cell division. Shortened chromosomes limit the number of times a cell can divide. When the telomere is shortened to a certain extent, important genetic material is lost if the cell continues to replicate DNA. At this point, normal cells will enter the cell aging process, or a growth arrest will occur. In both cases, the cells no longer continue to divide.
However, cancer cells have the ability to continue to replicate their DNA without entering the aging phase. In many cancers, this unrestricted cell division ability is due to the production of a telomerase . This enzyme maintains the end of the chromosome, preventing it from becoming shorter. Telomerase is a normal protein that is present during fetal development. However, in most cells of an adult, this enzyme no longer exists because the gene for this enzyme is no longer expressed (transcribed and translated). But in some cancer cells, the genes encoded by this enzyme are reactivated, eventually leading to uncontrolled replication.
The following animation shows two chromosomes, with active telomerase on the right and inactive telomerase on the left.
In cancer cells, chromosomes are not shortened even in the absence of telomerase activity, a phenomenon that can be explained by other mechanisms. Because only the length of the telomere is maintained, cell division is not restricted. hTERT is a gene encoded by the active component of telomerase and is considered to be a proto-oncogene because its abnormal expression can lead to uncontrolled cell growth.
BCL -2
BCL-2 (ie, B cell lymphoma gene 2) protein is closely related to cell membrane and cell membrane activity. The BCLl-2 protein is a component of a complex signaling system that regulates apoptosis. Apoptosis can be induced by a variety of signals, including irreparable damage to DNA. This cell prevents further expansion of cell damage in a suicidal manner. Bcl-2 prevents apoptosis. Therefore, its overexpression prevents the apoptosis of damaged cells, which leads to the continuous division of the mutant cell line, which eventually leads to the occurrence of cancer. In addition, overexpression of BCL-2 can also lead to the metastasis of some cancers.
If the control of apoptosis is disrupted, the anticancer drugs that induce apoptosis will not be as effective as before. So some drugs are being developed to inhibit the expression of the BCL-2 gene, making other anticancer drugs more effective (and less dose). One such drug is Genasense, which is an antisense nucleotide. This drug can reduce the production of BCL-2 protein in the first phase of the experiment, and is currently undergoing Phase II and Phase III trials, which can be used as adjunctive drugs for the treatment of various malignant tumors. on antisense drugs more information (antisense drugs) in.
In addition, there are some drugs that indirectly reduce the BCL-2 protein content, such as all-trans retinoic acid, paclitaxel, vincristine, and docetaxel. These drugs are often used in combination with other chemotherapy drugs. There are also some new drugs (not yet tested in humans), such as “BCL-2 binding polypeptide” can cause BCL-2 protein inactivation, antimycin A can bind to “Bcl-2 related protein”.
Since overexpression of the BCL-2 gene affects the success rate of cancer treatment, determining whether it is functioning properly becomes a valuable diagnostic basis. This proto-oncogene can cause gene overexpression by translocation, which in turn activates into an oncogene. High levels of BCL-2 protein are found in many different cancers.
EGFR
The epidermal growth factor receptor (EGFR) is a transmembrane protein. It is partially outside the cell and then passes through the cell surface, extending a portion to the cytoplasm. Its function is as a cellular antenna. When a specific protein binds to EGFR, the two epidermal growth factor receptor proteins bind to each other. Examples of EGFR are: epidermal growth factor or transforming growth factor alpha. The two proteins that are bound together are called dimers, and the formation of dimers activates the receptors, leading to autophosphorylation. These two proteins add a small molecule of chemicals to each other called a phosphate group. Activation of EGFR can increase cell proliferation and survival activity.
In many cancers, the EGFR gene is mutated or overactive. These cancers include breast cancer, lung cancer, esophageal cancer, and head and neck cancer. In cancer cells and other cells present in tumors, excessive activity of EGFR leads to blood vessel growth, excessive cell division, enhanced cell survival and cell movement, leading to the spread of cancer.
Currently targeted therapies for EGFR mutations are: monoclonal antibodies and tyrosine kinase inhibitors (TKI). Some antibodies, such as rituximab (Erbitux ® ) and panitumumab (Vectibix®), bind to the extracellular portion of EGFR to prevent ligand-activated receptors. Some TKIs, such as gefitinib (Iressa®) and erlotinib (Tarceva ® ), bind to the intracellular portion of EGFR to prevent activation.
Oncogene list
| cancer Gen | Function / activation | cancer* | references |
| ABL | Promotes cell growth through tyrosine kinase activity | Chronic myeloid leukemia | 17 18 |
| AF4/HRX | Fusion affects HRX transcription factor/methyltransferase. HRX is also known as MLL, ALL1 and HTRX1 | Acute leukemia | 19 |
| AKT-2 | Coding for serine/threonine protein kinase | Ovarian cancer | 19 20 |
| ALK | Encoding for tyrosine kinase receptor | Lymphoma | twenty one |
| ALK/NPM | Translocation produces fusion protein and nuclear phosphoprotein (npm) | Large cell lymphoma | twenty two |
| AML1 | Coding for a transcription factor | Acute myeloid leukemia | 19 |
| AML1/MTG8 | New fusion protein produced by translocation | Acute leukemia | 19 |
| AXL | Encoding for tyrosine kinase receptor | Hematopoietic malignant disease | 19 |
| BCL -2, 3, 6 | Blocking apoptosis (programmed cell death) | B-cell lymphoma, leukemia | 18 19 |
| BCR/ABL | New proteins produced by fusion of bcr and abl trigger unregulated cell growth | Chronic myeloid leukemia, acute lymphoblastic leukemia | 23 18 |
| C- MYC | Transcription factors that promote cell proliferation and DNA synthesis | Leukemia, breast cancer, stomach cancer, lung cancer, cervical cancer, colon cancer, neuroblastoma, glioblastoma | twenty four |
| DBL | Guanine nucleotide exchange factor | Diffuse B-cell lymphoma | 19 |
| DEK/CAN | Fusion produces new proteins | Acute myeloid leukemia | 19 |
| E2A/PBX1 | Fusion produces new proteins | Acute B cell leukemia | 19 |
| EGFR | Cell surface receptors that trigger cell growth by tyrosine kinase activity | Squamous cell carcinoma | 23 18 |
| ENL/HRX | Fusion protein produced by translocation t(11; 19) | Acute leukemia | 19 |
| ERG/TLS | The fusion protein is produced by the t(16:21) translocation, and the erg protein is a transcription factor. | Myeloid leukemia | 18 19 |
| ERBB | Cell surface receptors that trigger cell growth by tyrosine kinase activity | Glioblastoma, squamous cell carcinoma | twenty three |
| ERBB-2 | Cell surface receptors that trigger cell growth through tyrosine kinase activity; also known as HER2 or neu | Breast cancer, salivary gland cancer, ovarian cancer | 23 20 |
| ETS -1 | Transcription factor | Lymphoma | 25 |
| EWS/FLI- 1 | Fusion protein produced by translocation t (11:22) | Ewing sarcoma | 18 26 |
| FMS | Tyrosine kinase | sarcoma | 27 |
| FOS | API transcription factor | Osteosarcoma | 18 7 |
| FPS | Tyrosine kinase | sarcoma | 27 |
| GLI | Transcription factor | Glioblastoma | 28 |
| GSP | Membrane-associated G protein | Thyroid cancer | 7 |
| HER2 / neu | Overexpression of conductive kinase due to gene amplification | Breast cancer, cervical cancer | 23 7 |
| HOX11 | Transcription factor | Acute T-cell leukemia | 19 |
| HST | Fibroblast growth factor | Breast cancer, squamous cell carcinoma | 19 |
| IL-3 | Cell signaling molecule | Acute precursor B-cell leukemia | 19 |
| INT-2 | Fibroblast growth factor | Breast cancer, squamous cell carcinoma | 19 |
| JUN | API transcription factor | sarcoma | 20 7 |
| KIT | Tyrosine kinase | sarcoma | 20 7 |
| KS3 | Herpesvirus-encoded growth factor | Kaposi’s sarcoma | 7 |
| K- SAM | Fibroblast growth factor receptor | Gastric cancer | 19 |
| LBC | Guanine nucleotide exchange factor | Myeloid leukemia | 7 29 |
| LCK | Tyrosine kinase | T-cell lymphoma | 7 |
| LMO1 , LMO2 | Transcription factor | T-cell lymphoma | 7 |
| L- MYC | Transcription factor | Lung cancer | 19 20 |
| LYL -1 | Transcription factor | Acute T-cell leukemia | 19 |
| LYT -10 | Transcription factor. Also known as NFκB2 | B-cell lymphoma | 7 |
| LYT -10/ Cα1 | Transfer of the fusion protein immediately adjacent to the Cα1 immunoglobulin locus by (10; 14) (q24; q32) lyt-10 | 19 | |
| MAS | Angiotensin receptor | Breast cancer | 7 |
| MDM -2 | Encoding a protein that inhibits and degrades p53 | sarcoma | 19 20 |
| MLL | Transcription factor/methyltransferase (also known as hrx and ALL1) | Acute myeloid leukemia | 18 7 |
| MOS | Serine/threonine kinase | Lung cancer | 19 30 |
| MTG8/AML1 | Fusion of transcriptional repressors with transcription factors. AML1 is also known as RUNX1. | Acute leukemia | 19 |
| MYB | Transcription factor | Colon cancer, leukemia | 19 |
| MYH11/CBFB | Generation of new proteins by transcription factor fusion (through inversion on chromosome 16) | Acute myeloid leukemia | 19 |
| NEU | Tyrosine kinase. Also known as erbB-2 or HER2 | Glioblastoma and squamous cell carcinoma | 23 19 |
| N- MYC | Cell proliferation and DNA synthesis | Neuroblastoma, retinoblastoma, lung cancer | 23 19 |
| OST | Guanine nucleotide exchange factor | Osteosarcoma | 7 |
| PAX -5 | Transcription factor | Lymphocyte-like B-cell lymphoma | 7 |
| PBX1/E2A | The fusion protein was formed by t (1:19) translocation.Transcription factor | Acute precursor B-cell leukemia | 19 |
| PIM -1 | Serine/threonine kinase | T-cell lymphoma | 7 |
| PRAD -1 | It encodes for cyclin D1 and is involved in cell cycle regulation. | Breast cancer, squamous cell carcinoma | 19 |
| RAF | Serine/threonine kinase | Multiple cancer types | 19 |
| RAR / PML | The fusion protein was formed by t (15:17) translocation.Retinoic acid receptor. | Acute promyelocytic leukemia | 23 19 |
| RAS -H | G protein. Signal Transduction. | Bladder Cancer | 19 |
| RAS -K | G protein. Signal Transduction. | Lung cancer, ovarian cancer, bladder cancer | 19 20 |
| RAS -N | G protein. Signal Transduction. | Breast cancer | 19 |
| REL / NRG | A fusion protein formed by deletion of chromosome 2.Transcription factor | B-cell lymphoma | 19 20 |
| RET | Cell surface receptors. Tyrosine kinase | Thyroid cancer, multiple endocrine neoplasia type 2 | 23 19 |
| RHOM1 , RHOM2 | Transcription factor | Acute T-cell leukemia | 19 |
| ROS | Tyrosine kinase | sarcoma | 7 |
| SKI | Transcription factor | cancer | 7 |
| SIS | Growth factor | Glioma, fibrosarcoma | 7 |
| SET / CAN | A fusion protein formed by rearrangement of chromosome 9. Protein localization | Acute myeloid leukemia | 7 31 |
| SRC | Tyrosine kinase | sarcoma | 19 32 |
| TAL1 , TAL2 | Transcription factor. TAL1 is also known as SCL | Acute T-cell leukemia | 19 33 |
| TAN -1 | Alteration of Notch cell receptor formation by t(7:9) translocation | Acute T-cell leukemia | 19 34 |
| TIAM1 | Guanine nucleotide exchange factor | T-cell lymphoma | 7 35 |
| TSC2 | GTPase activator | Kidney cancer, brain tumor | 7 36 |
| TRK | Receptor tyrosine kinase | Colon cancer, thyroid cancer | 19 37 |
*The type of cancer listed in the above table is the main type of cancer associated with each oncogene, but this is not an exhaustive list.
For more information, visit the Cancer Genome Anatomy Project .
Summary of this section: Oncogene
cancer Gen
- An oncogene is a mutant form of a normal cellular gene (proto-oncogene).
- The protein product of the proto-oncogene can stimulate cell division and/or inhibit cell death.
- The proto-oncogene can be likened to the accelerator pedal of a car.
- Normally, internal and external signals strictly control the activity of proto-oncogenes, but oncogenes are defective, so when they do not receive the appropriate signal, they are also in the “on” mode.
- Oncogenes also help cells ignore negative signals that prevent healthy cell division.
- Oncogenes can cause cells to continue to divide, even if they do not receive any growth-promoting signals.
- The different cellular effects of some known oncogenes are listed below:
- HER-2/neu
- HER-2/ neu encodes a cell surface receptor that stimulates cell division.
- The HER-2/ neu gene can be amplified by up to 30% in human breast cancer.
- RAS
- The Ras gene product is involved in the kinase signaling pathway, ultimately controlling gene transcription, regulating cell growth and differentiation.
- Overexpression and expansion of RAS can result in sustained cell proliferation.
- MYC
- Myc protein is a transcription factor that controls the expression of some genes.
- Myc has been shown to be involved in avoiding cell death mechanisms.
- MYC oncogenes may be activated by gene rearrangements or amplification.
- SRC
- SRC is the first gene discovered to date.
- The Src protein is a tyrosine kinase that regulates cellular activity.
- hTERT
- hTERT encodes an enzyme that maintains the end of a chromosome, called telomerase.
- In most normal cells, telomerase is only present during fetal development.
- In adult cells, the activation of hTERT gives them the ability to divide indefinitely.
- hTERT encodes an enzyme that maintains the end of a chromosome, called telomerase.
- BCL-2
- Bcl-2 protein acts to prevent cell death (apoptosis).
- Overexpression of BCL-2 results in the continued division of mutant cells.
- HER-2/neu
Understanding the process: oncogenes
“Understanding the Process” is an educational game that tests your knowledge. Start the game:
- Select the appropriate options from the right column and drag them into the corresponding boxes on the left. Please note that you will only use five of the six options to complete the game.
- When you’re done, click “Query” to see your correct rate.
- For incorrect answers, click on “Description” to review the information.
- If you want to try again, please select “Reset” and start the game again.
Flow chart fill in the blank: oncogene
Sequential process
-
1
-
2
-
3
-
4
-
5
Biological process
-
Apoptosis
-
Tumor formation
-
Abnormal cell division
-
Oncogene produces abnormal protein
-
Proto-oncogene mutation
-
Proto-oncogene undergoes chemical mutagenesis
The figure below lists the functions of some of the above oncogenes. Many other oncogenes have these similar activities.

Tumor suppressor
Tumor suppressor genes act on many key cellular processes such as transcription, DNA repair, and cell-to-cell communication. Loss of function of these genes will lead to abnormalities in the way cells behave.
To continue the previous analogy, the tumor suppressor gene is like the brake system of a car. If you think that every copy of the tumor suppressor gene acts on the “brake” of the cell, then this metaphor is convincing. When both copies of the tumor suppressor gene are active (indicated by the highlighted gene and the stop sign in the image below), the cell stops dividing (like the car will stop driving).
When a set of copies of a tumor suppressor gene is defective, the cell has another set of copies that are working. Just like the car parking, only the front or rear brakes are used instead of the front and rear brakes. Although the parking effect is not so good, it can still stop. Therefore, when a cell has a copy of a defective tumor suppressor gene, the cell can still control its cell division. But if another set of tumor suppressor copies does not work, the cells lose control of cell division.
In the following sections, we will introduce the retinoblastoma ( retinoblastoma , Rb) tumor suppressor gene, it would be a good example to illustrate this mechanism.
All cancers show alterations in one or more tumor suppressor genes and oncogenes . In normal cells, the proteins produced by the two together control cell division, and in cancer cells, such control is abnormal.
Since these genes are important in the development of cancer, some specific tumor suppressor genes and related cancers will be described in detail in the following sections. The genes involved in these processes are increasing almost every day, but here we can only introduce some of the genes that we know so far, and we can understand how cancer causes imbalance in cell function.
Introduction to p53
The p53 (or TP53 ) gene was discovered in 1979 and is one of the most closely related genes to date. The gene is located on chromosome 17, and its protein product functions as a transcription factor. The gene controlled by p53 protein is involved in cell division and viability. Like other tumor suppressor genes, p53 acts to prevent uncontrolled cell growth.
The p53 protein interacts directly with DNA and interacts with other proteins that govern cell behavior. When DNA is found to be damaged or other cells are damaged, p53 immediately initiates a mechanism of cell death (ie, apoptosis). The p53 protein plays a key role in maintaining normal cell regulation, especially in about half of tumors, regardless of its type or origin, and defects in the p53 gene are found. In a person’s life, mutations (sporadic mutations) that cause the loss of p53 gene activity may occur or this type of mutation is obtained by inheritance.
Discovery of the p53 gene
In 1979, scientists discovered a new protein. This protein binds to a transformin-large T antigen in simian virus 40 (SV40). This protein is more common in virally transformed cells (with immortality and carcinogenicity) compared to normal cells. The protein and its corresponding gene were named p53, which was named according to the mass of the protein being 53 kilodaltons. The p53 gene is located at position p13 of chromosome 17.
Although p53 is the second tumor suppressor found after Rb, scientists did not understand its role in cells until 10 years after p53 discovery. Because of the increased content of p53 in transformed cells, researchers initially thought it would act like an oncogene.Theinitial findings also support this view. Scientists have discovered that when the p53 gene enters the cell, the cell transforms. But the researchers later discovered that the p53 gene that enters the cell is actually a mutant of this gene, because the normal function of the p53 gene is to prevent cell transformation!
There is a variety of evidence that the p53 gene is a tumor suppressor. Since 1989, the p53 research has made continuous progress. The p53 protein is involved in multiple cellular processes. However, the clinical use of p53, such as the feasibility of identifying cancer cells, remains questionable. The focus of current research is to test and evaluate the feasibility of p53 clinical treatment to determine whether it can repair the p53 gene or replace the damaged p53 gene.
Function of p53 protein
The p53 protein plays an integral role in the cell. Normally, this protein is present in all types of cells. This protein is located in the nucleus and functions as a transcription factor. The p53 protein is located in the center of a large protein network, “monitoring” the health of cells and cellular DNA. The p53 protein acts like a well-organized, coordinated “orchestral conductor” that detects and controls any cell damage in time. When damage is detected, the function of the p53 protein is to help determine whether to repair damaged cells or induce damage to cells (ie, apoptosis).
As a transcription factor, p53 promotes transcription of a set of target genes. Of these genes, p21 is the most important gene. The product of the p21 gene is a negative regulator of cytokine-dependent kinase. This enzyme plays a key role in the normal functioning of the cell cycle and in the final cell division. By promoting the transcription of the p21 gene, p53 protein prevents cell proliferation. This blocking action gives the cell a chance to repair. In the event of severe DNA damage, p53 protein can help initiate cell death. The death of severely damaged DNA cells is beneficial to the body because it prevents the continued replication of harmful mutant cells.
As described in this section, all cancer cells contain a combined mutation of a tumor suppressor gene and an oncogene. Removal of functional p53 (the “guard” of the genome) from the cell leads to the accumulation of more DNA damage, and the continued division of more damaged cells.
Mutations in the p53 gene are the most common genetic alterations in cancer cells. This mutation occurs in addition to the growth and development of the individual (sporadic mutations), and some mutations occur in certain types of malignancies that are involved in the inheritance of impaired p53 replicas. For example, the Li-Fraumeni familial tumor syndrome is often associated with a variety of malignant tumors. In addition, there are several viruses have evolved and have the inactivation of the p53 protein pathway.
Since this protein plays a central role in regulating cell division, a large amount of research is devoted to developing a safe method for restoring the function of the p53 gene.
In 2014, the researchers discovered another form of transcriptional splicing of the p53 gene that promotes cancer development and spread. The new form is called p53Ψ. This work was carried out in cell culture, mouse antibodies, and rabbit antibodies.
Abnormality of p53 protein and cancer development
A cell lacking functional p53 may or may not develop cancer. Accordingly, a cell with normal p53 function may eventually lead to the formation of cancer cells. As discussed in the previous section of Gene Mutation, a cell’s DNA must undergo some changes before mutation can occur. One of the functions of p53 is to monitor the state of cellular DNA. P53 works with a range of other proteins to identify and repair damaged DNA. The cellular responses stimulated by damaged DNA are: repair, stop cell division and cell death. Damage to the p53 gene does increase the likelihood of cancer development. Keep in mind that since the p53 gene is a tumor suppressor gene, only two copies of the gene are inactivated to have an effect. Here are a few ways to inactivate p53:
Gene mutation
Changes in the p53 gene have different effects on gene activity, depending on where the gene is altered.
- Mutations may occur in regulatory regions. These regions of the gene control the frequency and timing of gene transcription, and this region is called initiation. Mutations that occur in the promoter region can result in decreased or lost p53 in the cell.
- Mutations may occur in the protein coding region of a gene. This mutation has several ways to influence gene expression (or protein activity):
- The activity of the transcription factor p53 is reduced. This will affect the expression of the p53 target gene. Target genes for p53 include p21 (a protein involved in cell cycle regulation), Bax (a protein that induces apoptosis), and thrombospondin 1 (an inhibitor of angiogenesis).
- Changes in the p53 protein make it more susceptible to degradation. If the p53 proteins in the cells are degraded faster than normal, they will no longer have normal tumor suppressor function.
Virus inactivation
One of the functions of p53 is to “guard” the genome. Viral infection introduces foreign DNA into the cell, and p53, along with other proteins, is responsible for reacting to foreign DNA in the cell, such as stopping cell division and stimulating cell death. However, several viruses have evolved methods to try to inactivate p53 proteins to avoid these reactions. An example of this is simian virus 40 (SV40). When the cells are infected with SV40, viral proteins are produced in the cytoplasm. One of the proteins produced is called the large T antigen. The function of this protein is to bind to the p53 protein and inactivate it. Other viruses, such as hepatitis and human papillomavirus, also produce similar proteins.
If the cell loses its normal working p53, the cell will continue to divide, even if there is DNA damage. In the absence of p53, the instability of the gene is reflected in the increased likelihood of genetic mutations and aneuploidy. Increased genetic damage results in the accumulation of defective tumor suppressor genes and oncogenes.
Introduction to the retinoblastoma gene Rb
The retinoblastoma ( Rb ) gene can be a protein encoding a transcription factor. Rb proteins can indirectly regulate gene expression through interaction with transcription factors. In addition to this function, the Rb protein and several proteins that are closely related to it have several other functions, although there is not much evidence. Rb proteins and these proteins closely related to it are involved in the regulation of cell division.
Mutations in the Rb gene occur in a variety of malignancies. One of the most studied tumors in this area is retinoblastoma, and the Rb gene is named after the name of this ocular tumor. The disease is common in young children. Two different types have been identified:
- Scattered hair: Can affect anyone, is the disease caused by genetic changes (mutations) that occur in the patient’s life.
- Familial type: The patient obtains a copy of the defective gene from her parents by inheritance. The gene copy in each cell of the patient has a normal and one defect.
Like other tumor suppressor genes, the tumor’s genetic phenotype is less pronounced unless both copies of the gene are damaged. Although it is not possible to have a mutation in the Rb gene in any cell, secondary mutations in the gene may occur in many cells of our body, including the eye. Familial patients may have other forms of tumors, especially osteosarcoma. Other tumors associated with mutations in the Rb gene are: lung cancer, breast cancer, and bladder cancer.
Detailed introduction of Rb gene and cancer
The Rb gene was originally found in a familial genetic form of retinoblastoma. This type of cancer primarily affects the eye and is most common in children. In the familial form of this gene, the patient’s cells already have a mutated Rb gene, so only a mutation in another Rb gene is required to make the cell lack a functional Rb protein. Although mutations are not common, and the probability of mutations in a particular gene is quite small, there are a large number of cells in our body that make this small probability of a genetic mutation occurring in at least a few cells. If these cells can grow in an uncontrolled way, it can cause cancer. Therefore, patients with familial retinoblastoma usually have multiple tumors. In the case of sporadic retinoblastoma, the patient’s cells have two functional gene copies of the Rb gene, so loss of Rb function requires simultaneous two mutations in the same cell . Therefore, patients with sporadic retinoblastoma usually develop only one tumor. In addition, patients with familial retinoblastoma are more likely to have tumor recurrence.
The researchers found that in patients with familial retinoblastoma, their osteosarcoma lost Rb activity. Osteosarcoma means that if a cancer metastasizes, nearly half of the patients may have a familial genetic form of the disease. In addition, the chances of women getting breast cancer are also related to Rb function. Rb is usually able to regulate the G1 cell cycle checkpoint, and studies have shown that some breast cancers are dysregulated at this checkpoint. This means that Rb is a contributing factor to the disease in this case. In addition, Rb also promotes other cancers, such as small cells and non-small cell lung cancer.
Rb function
The Rb gene plays an important role in maintaining the normal function of the cell cycle. Cells respond differently to various signals in their environment that regulate their growth, division, dormancy, and apoptosis. If these signals are destroyed, it can lead to uncontrolled cell growth, which ultimately leads to the formation of cancer. The regulation of cell division processes involves the overall role of different signals.
The Rb gene product (pRb) has a growth inhibiting effect under normal conditions. It binds to transcription factors and thus inhibits transcription factors. Therefore, the Rb protein can indirectly control the expression of many genes. Some of these genes produce proteins that drive cell division. Therefore, Rb protein can act to slow or prevent cell division. regulatory proteins (e.g. Rb protein) will change and the whole cell body to produce a significant effect. In addition to regulating the cell cycle, Rb protein also has an effect on apoptosis. Apoptosis is a very important cellular function that enables damaged cells to enter a programmed death process. If the mutation in the cell is irreparable, the cell will be cleared by apoptosis. This mechanism also helps to clear cells that have uncontrolled growth potential and eventually grow into cancer cells. Any change in cellular function that weakens or loses the mechanism of apoptosis may cause damage to the entire cell population. Several different types of genetic damage can inactivate the Rbgene. Mutations that completely lose the function of the Rb protein (ie, null mutations) are common in cells that lose the functional Rb protein.
In fact, the loss of Rb may in some cases help patients respond to chemotherapy. Studies have shown that breast cancer patients who have a lack of Rb function will respond better to neoadjuvant chemotherapy (neoadjuvant chemotherapy is the chemotherapy they received before surgery).
Detailed introduction of Rb function
Recent studies have shown that in addition to the function of regulating cell growth and apoptosis, Rb-related proteins have different activities depending on the stage of the cell cycle and the location of these proteins in the nucleus. In addition, some studies have shown that proteins like Rb regulate the transcription of ribosomal RNA and transcript RNA. That is to say, pRb can control the “transcription” process and the “post-transcriptional” process in cells.
In addition to acting as a transcription factor, pRb may also have tumor suppressive effects. Rb proteins have been shown to be associated with chromatin-modifying proteins such as histone deacetylase (HDAC). It is believed that histone deacetylases affect the transcription process by removing acetyl groups from histones. This modification results in a tighter relationship between DNA and nucleosomes. The stronger interaction of DNA with histones makes it more difficult to bind transcription factors such as E2F to its target sites within DNA. Although the role of Rb in this process remains unclear, studies have shown that at least some of the histone deacetylases do not function properly in the absence of Rb.
Introduction to APC
Mutations in the APC (adenomatous polyposis) gene are closely related to hereditary and sporadic colon cancer. As we will describe in the next section, APC proteins, like many tumor suppressor proteins, control gene expression that plays a key role in cell division.
Most colon cancer cases are diagnosed as slow-moving diseases that can last for years. On chromosome 5, inactivation of the APC gene is thought to be the main cause of accelerated cell proliferation and colonic polyps. Several genetic changes occur during the conversion of normal colon cells into potential tumor “seed” cells. In many cases, mutations in the APC gene are thought to be the earliest changes. This statement can be indirectly demonstrated by the detection of patients who have acquired the APC mutant gene by heredity . These patients have familial adenomatous polyposis, and the patient’s colon is full of polyps. Every polyp has the potential to develop into cancer, so individuals with genetic mutations are much more likely to develop cancer. This is very similar to the genetic form of retinoblastoma described in the ” Rb Gene” section. The difference is that this type of patient does not require two somatic mutations in the cell to cause APC to lose function, but a genetic mutation in any cell can cause major problems.
By comparing cell mutations at different stages of cancer, a possible sequence of genetic mutations in a part of colon cancer can be established. In this sequential model, the APC gene mutation occurs in the first step, producing highly proliferating cells that form polyps and eventually develop into cancer.
APC function
Since the lack of functional APC protein can lead to an increase in cell division, normal functioning APC proteins can inhibit cell division in some way. The formation of a complex of APC protein with β -cateninleads to the degradation of β-catenin (β-catenin is a transcription factor). In the absence of APC protein, excess β-catenin will accumulate in the nucleus. Beta-catenin binds to another protein in the nucleus to form a complex that binds to DNA and initiates transcription of several genes. One target gene in this complex is called c-myc , a known oncogene . C-myc itself is a transcription factor for several genes that controls cell growth and division. Thus, mutations in the APC gene can lead to a series of chain reactions that ultimately accelerate cell division. An illustration of the APC features is shown below.
Of course, many other factors may also affect the expression of genes and their products, but the APCgene mutations appear to [beta] -catenin and c-myc much related, [beta] -catenin and c-myc increased number of proliferating cells may lead to Speed up.
Studies have shown that the addition of APC protein to colon cancer cells lacking APC protein has the function of reducing tumor cell growth. The decline in growth is due to increased apoptosis, which means that APC has the function of controlling cell death and growth. Therefore, the loss of APC gene has a certain effect on regulating the balance between cell growth and cell death, so the APC gene controls the number of cells.
Tumor suppressor: BRCA
BRCA proteins have multiple functions. One of the important functions is to repair DNA damage. This is also involved in the regulation of gene expression. The BRCA -1 gene is involved in the activation of another tumor suppressor gene, p53 , whose target gene is p21 .
BRCA proteins also interact with transcription factors and other transcriptional components to control the activity of several other genes. When the BRCA gene is inactivated, DNA repair and gene regulation are affected. Intensification of DNA damage produces cells that cluster key mutations, leading to the formation of malignant cells. Cells lacking the BRCA gene often exhibit chromosomal breaks, severe aneuploidy, and centrosome amplification.
At the molecular level, the structure of the BRCA -1 gene and the BRCA -2 gene explain their susceptibility to mutations. They contain a high proportion of repetitive DNA, which is very rare in human genes. This highly repetitive DNA can lead to instability and rearrangement of the genome.
A number of studies have confirmed that the lack of BRCA gene products is closely related to the occurrence of sporadic or hereditary malignancies.
Detailed introduction of BRCA gene and ovarian cancer
The BRCA gene is named for its association with breast cancer, and mutations in these genes are also associated with ovarian cancer. The genetic pattern and distribution of ovarian cancer are similar to those of breast cancer, but there are some differences. Hereditary ovarian cancer has many histological abnormalities, which are “medium and poorly differentiated” and have invasive growth characteristics, usually found in the late stage. In addition, BRCA mutant gene carriers have a higher incidence of fallopian tube lesions. Ovarian tumors with benign or low-grade malignant potential should not be considered as pre-invasive ovarian cancer, regardless of whether the patient is a mutated gene carrier.
BRCA features
Mutations in BRCA-1 and BRCA-2 are associated with some breast and ovarian cancers. These two genes have different functions in the cell. As with the previously described tumor suppressor genes, mutations in these two genes can occur naturally or genetically. Individuals genetically acquiring mutations in the BRCA-1 or BRCA-2 gene are more likely to develop breast cancer. Carriers of BRCA mutations (if they live up to 85 years of age) have an 80% chance of developing breast cancer in their lifetime. Carriers of the BRCA-2mutation have a 10-20% chance of developing ovarian cancer in their lifetime, while carriers of the BRCA-1mutation have a 40-60% chance of developing ovarian cancer. The presence of these mutations also increases the likelihood of prostate cancer, pancreatic cancer, colon cancer, and other cancers. For any individual, the total likelihood of illness depends on individual genetic and environmental risk factors. 5-10% of breast cancer cases are thought to be associated with mutations in BRCA-1 and BRCA-2 genes.
BRCA gene and estrogen ( Estrogen )
BRCA gene mutations in certain organs and tissues (such as breast, ovarian) cancer related. This means that estrogen may play a role in the cancer development of these tissues. Fluctuations in estrogen levels (such as during puberty, menstruation, pregnancy, and menopause) are associated with cancer development. During puberty and pregnancy, elevated estrogen levels can lead to the proliferation of mammary epithelial cells, which requires these cells to have a greater ability to repair DNA. If cell proliferation (ie cell division) is accompanied by a decrease in the efficiency of DNA repair, it can lead to the formation of cancer. In the animation below, the pink dots indicate estrogen. And estrogen can stimulate cell division and produce cancer cells.
If a copy of the BRCA gene is mutated such that the only functional gene does not function, DNA repair defects can occur. If both copies of the repair gene lose function, the likelihood of the cell gaining a mutation increases, leading to tumorigenesis. When a person inherits a defective BRCA gene copy, all of his cells will contain this mutation. At this time, if another replica of the BRCA gene in any one cell is mutated, DNA repair may be difficult. If an individual does not have a BRCA allele due to heredity , then two separate mutations are required to cause cancer, and the two sporadic mutations must occur in the same cell. However, the possibility of two simultaneous mutations in the same cell is minimal, which explains why cancer usually occurs when people are older.
For more information on BRCA and estrogen, please refer to Robert A. Weinberg, Sections 3, 4, and 7 of Cancer Biology .
Details of BRCA gene and survival rate
There have been a number of studies to determine the difference in survival between BRCA mutant gene carriers and sporadic tumor patients, but the results obtained are not consistent. It may be due to different experimental design schemes or other reasons, such as improper matching between the control (sporadic tumor patients) and the tester (mutant gene carriers). However, according to the characteristics of the tumor they are suffering, the BRCA mutation gene carriers have a poor prognosis. However, compared with patients with sporadic tumors, the survival rate seems to be similar, probably because the tumor response to chemotherapy is “better”. The sensitivity of tumors to chemotherapy is increased, in part because of the high rate of tumor growth in patients. These tumors are sensitive to certain anticancer therapies such as gamma rays, cisplatin, and mitomycin C. These treatments can destroy DNA. The damaged DNA is normally repaired by a functional BRCA gene product. If the BRCA gene activity disappears, the cells cannot repair DNA as efficiently as normal, and the cells eventually die. The non-cancerous cells of the BRCAmutant gene carrier still have a functional BRCA gene that can repair damaged DNA.
Tumor suppressor gene list
| Tumor suppressor | Features | Cancer * | references |
| APC | Control the function of specific transcription factors. These transcription factors are involved in the oncogenesis, development and homeostasis of some cells, including epithelial cells, lymphocytes, and the like.
APC is also involved in cell proliferation and other cellular activities such as migration and adhesion. |
Familial adenomatous and non-hereditary colorectal cancer | |
| BRCA1, BRCA2 | DNA damage repair | Inherited breast cancer; ovarian cancer | |
| CDKN2A | Locus encoding the tumor suppressor genes p16 and p14ARF | Brain tumor | 1 |
| DCC | Netrin-1 receptor.Regulates cell proliferation and intestinal epithelial cell apoptosis | Colorectal cancer | |
| DPC4 (SMAD4) | Transfer factors involved in development; affect metastasis and tumor invasion | Colorectal tumor, pancreatic tumor | |
| MADR2/JV18 (SMAD2) | Signals that mediate growth factor receptors.Assist in the transport of SMAD4 into the nucleus | Colorectal cancer | |
| MEN1 | For the encoding of the menin protein, the menin protein interacts with transcription factors, DNA repair proteins, cytoskeletal proteins, etc., but its function is not clearly defined. | Multiple endocrine neoplasia type 1 | |
| MTS1 | Cyclin-dependent kinase inhibitors; regulate cell cycle from G1 to S phase. | Melanoma | |
| NF1 | GTPase activating protein (RAS-GAP) | Neurofibromatosis type 1 | |
| NF2 | ERM protein; the plasma membrane is organized by assembling protein complexes and attaching them to actin. | Neurofibromatosis type 2 | |
| P53 | Encoded for the transcription factor of p21, p21 is a protein that blocks the cell cycle in the G1 phase. P53 acts on signals that integrate cell size, DNA integrity and chromosome replication. | Bladder cancer, breast cancer, colorectal cancer, esophageal cancer, liver cancer, lung cancer, prostate cancer and ovarian cancer; brain tumor, sarcoma, lymphoma and leukemia | |
| PTEN | Lipid phosphatase regulates cell survival | Cowden’s syndrome; increased risk of breast and thyroid cancer | 2 |
| Rb | Binding and inhibiting E2F transcription factors. Stop cell cycle progression | Retinoblastoma, sarcoma; bladder cancer, breast cancer, esophageal cancer, prostate cancer and lung cancer | |
| VHL | Cell cycle regulation. Can increase the stability and activity of p53 | Renal cell carcinoma | 1 |
| WRN | DNA helicase and exonuclease. Participate in repairing DNA breaks. | Werner syndrome | 2 |
| WT1 | Transcription factor; plays an essential role in development | Wilms tumor (pediatric kidney cancer) | 1 |
* The types of cancer listed in the above table are the major cancer types associated with each tumor suppressor gene, but this is not an exhaustive list.
For more information, visit the Cancer Genome Anatomy Project .
1. “cancer genes”, author: Cooper G, Publisher: Jones and Bartlett Publishers, Publication Date: 1995.
2. “the genetic basis of human cancer,” the authors: Vogelstein B, Kinzler KW, Publisher: McGraw-Hill, Publication Date: 1998.
Summary of this section: Tumor suppressor gene
Tumor suppressor gene
- The protein product of a tumor suppressor gene can directly or indirectly prevent cell division or cause cell death.
- Tumor suppressor genes can be compared to the brake system of a car.
- Loss of function of a tumor suppressor gene can result in abnormal behavior of the cell.
- Here are some of the key tumor suppressor genes:
- P53
- A transcription factor that regulates gene control of cell division and cell death.
- It plays an important role in the response of cells to DNA damage.
- Helps make decisions between repair and induction of cell death.
- Rb
- Act to alter transcription factor activity.
- Cell division is controlled by acting as an inhibitor.
- APC
- APC proteins bind and stimulate the degradation of transcription factors.
- Loss of functional APC protein leads to an increase in cell division.
- BRCA
- BRCA proteins have a variety of functions, including repair of DNA damage and regulation of gene expression.
- Abnormal function of BRCA can lead to impaired DNA repair and gene regulation.
- P53
MicroRNAs
A gene is a long string of DNA that encodes RNA . For many years, scientists have been working on genes that encode messenger RNA (mRNA) because RNA is used to guide protein production. (See our overview of gene function .) Other RNAs, such as tRNA, snoRNA, and rRNA, are not used to directly produce proteins, but they indirectly help with protein production.
In 1993, scientists discovered a new type of RNA in a worm that is very short and surprisingly active. scientific experiments have shown that this RNA can regulate the activity of different genes. Although mRNA molecules can have thousands of nucleotides in length, this novel RNA has only a few tens of nucleotides. Within ten years, many other examples similar to this small RNA have been discovered. Scientific experiments have shown that these microRNAs (or miRNAs) can control a variety of different genes and cellular processes.
Another surprise comes from researchers looking for new miRNAs. Some of them come from their own genes, but many come from other genes, usually in non-coding regions, those that do not encode proteins. In addition, the produced miRNAs are not functional. They require several processing steps and ultimately complete their gene regulation activities with the protein.
Two different ways of producing functional miRNAs are shown in the figure below (illustrated from Wikimedia Commons ).
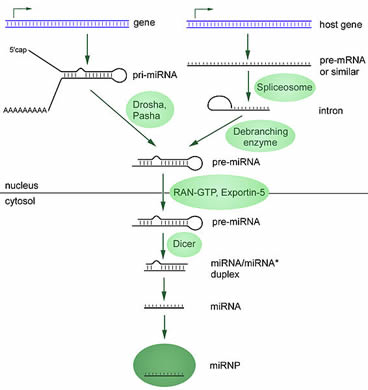
The final product of the “maturation” process is a short RNA that binds to a group of proteins (miRNP). miRNP is used to increase and decrease the activity of target genes. Mature miRNPs can bind to targeted mRNAs and prevent them from being used to produce proteins. They can also directly cause the destruction of targeted mRNA. Because in human cells, the activity of genes is tightly controlled to maintain balance, it is not surprising that miRNA production defects or activity defects are linked to several human diseases, including cancer. To learn more about the role of miRNAs in cancer, please read the following pages.
MicroRNAs and cancer
To date, microRNAs (miRNAs) are a very common RNA that regulates the effects of genes on a wide range of cellular activities. Changes in the activity of miRNAs can affect the activity of their target genes and lead to some visible changes, such as disease. Cancer is a genetic change that alters the activity of a gene, so it is obvious that changes in miRNA can affect the development and/or spread of cancer. In fact, the current research on miRNA has made great progress and has affected many fields of cancer biology, detection, diagnosis and treatment. Thefollowing describes how miRNAs affect cancer.
miRNAs and Cancer Prevention
Chemicals found in food can affect the activity of many genes, including those that encode miRNAs. Currently, scientists are actively exploring the effects of diet on miRNA activity to increase or decrease the risk of cancer.
miRNAs as oncogenes and tumor suppressor genes
Because miRNAs control the activity of genes, they may be oncogenes or tumor suppressor genes, depending on their effects on cell growth. Those miRNAs that slow down cell division or cause cell death are tumor suppressors. If such miRNAs are lost, cells increase the chances of division and survival; those that increase cell division or survival are oncogenes. There are now many examples showing that miRNAs can play a role in a variety of cancer types in these ways.
miRNAs as Drivers of Tumor Cell Metabolism
For years, scientists have found that cancer cells rely more on glycolysis to produce energy than normal cells. This is called the Warburg effect. This energy production pathway allows cancer cells to take in more sugar than other cells, and cancer cells may also be affected by blood sugar levels in the body. In addition, cancer cells produce more lactic acid through glycolysis, which can change the environment around the cells. These changes together can lead to the development of the disease. It has recently been discovered that miRNAs can cause the Warburg effect by affecting the activity of tumor suppressor genes such as p53and oncogenes such as HIF1A .
miRNAs as biomarkers for cancer detection and diagnosis
Biomarkers are substances that indirectly indicate the presence of a disease or underlying disease. For example, blood cholesterol levels are a biomarker to indicate cardiovascular health. Usually blood tests such as the PSA test are also used to check biomarkers. Because miRNAs are associated with cancer, researchers are currently looking for miRNAs that can serve as cancer biomarkers in blood or other tissues. They pointed out that these miRNAs can also be used as markers for drug resistance and for guiding treatment.
miRNAs as targets for cancer therapy
As regulators of cellular activity, miRNAs should serve as targets for cancer therapy. Since a single miRNA can control a large number of genes, drugs that target miRNAs prove to be very effective, and they can immediately shut down or open the entire pathway of the miRNA.
In the case of breast cancer, the growth and survival of many breast cancers depend on female hormone estrogen and progesterone. Based on this phenomenon, doctors use anti-hormone treatments such as tamoxifen, raloxifene and aromatase inhibitors. A 2012 study showed that progesterone can cause cancer cells to return to the same state as stem cells, making them more difficult to treat. This change in cellular behavior is due to inhibition of a panel of miRNAs known as the miRNA 29 family. Researchers are looking for ways to increase the activity of these miRNAs in cancer cells, thereby reversing the characteristics of stem cells in cancer cells.
miRNA as a driver of drug resistance
Cancer drugs can sometimes fight cancer effectively, but only at the beginning. Over time, the patient’s sensitivity to the drug is reduced. It is this drug resistance that makes it very difficult to treat cancer. Therefore, most cancer research is investigating the causes and ways of this resistance development. There is increasing evidence that miRNAs contribute to this change in drug resistance, and that the abnormal levels of many miRNAs are closely related to drug resistance. However, when one of the miRNAs returns to normal levels, the cancer will regain sensitivity to the cancer drug.
NASA funded a research project in Emory to review the role of miRNAs in cancer development. Learn more about the Emory NSCOR study.